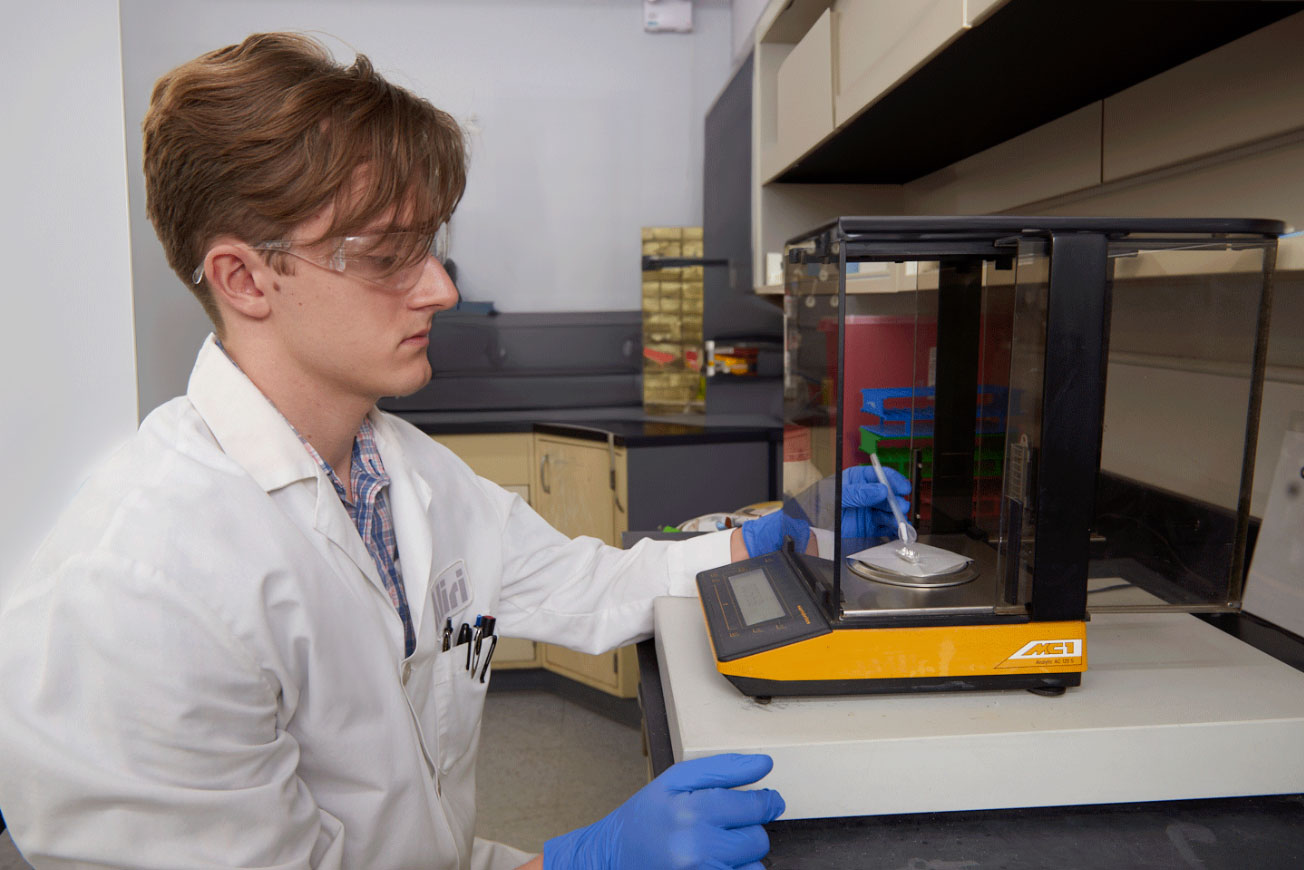
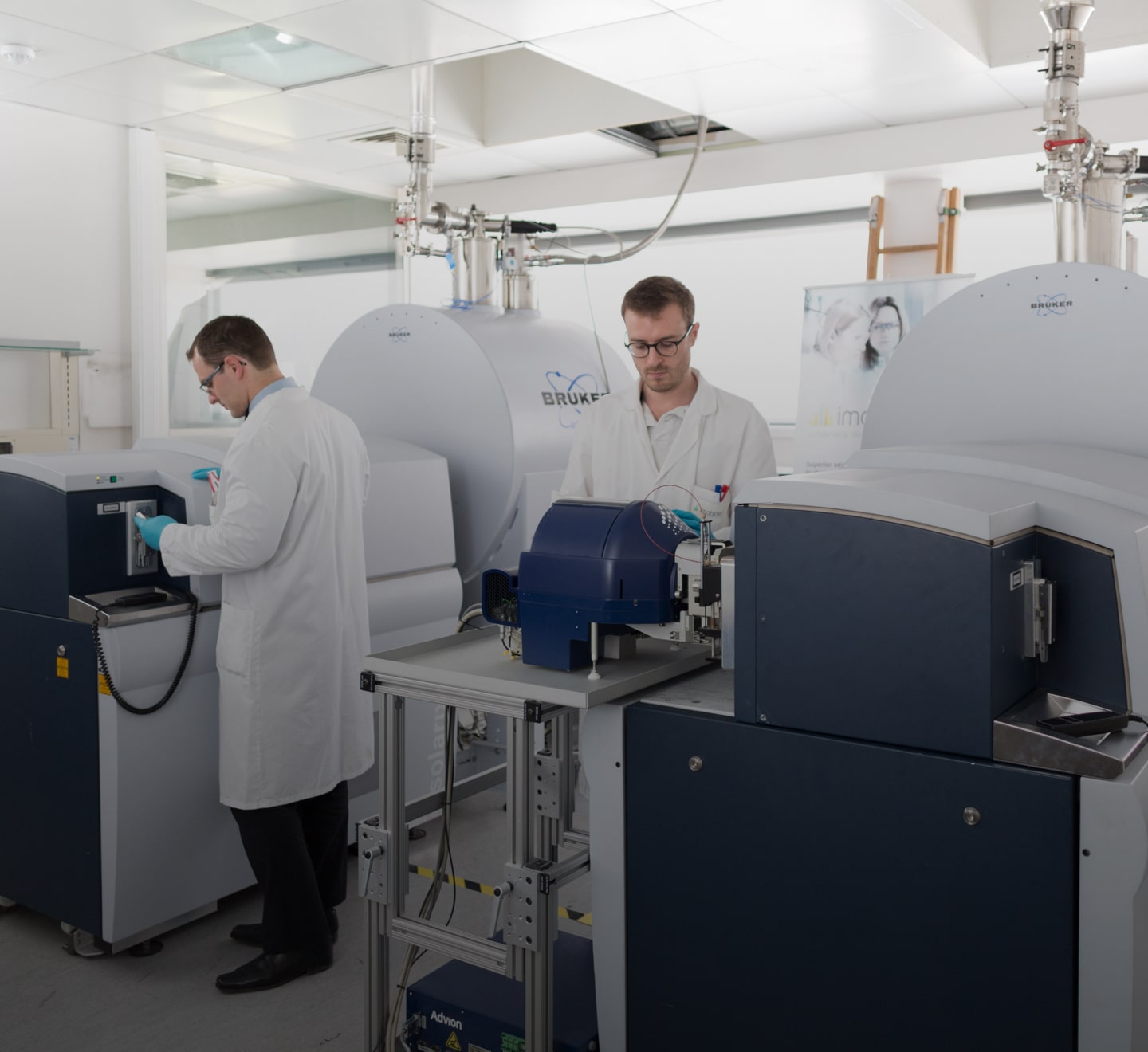
We provide a range of bioanalysis lab, spatial bioanalysis, and spatial biology solutions that will help you advance your molecule from discovery through clinical development and NDA approval, while being backed by a network of technical experts and seasoned scientists.
BioAnalysis
With patients’ lives on the line, you cannot afford any mistakes within your drug development program. That’s why finding the right bioanalytical lab to get you accurate and timely data will help you successfully file your IND with speed and confidence. We are committed to ensuring that your drug program is scientifically and regulatory compliant across all phases.
Read More
Spatial
Spatial bioanalysis and biology are solutions to help generate unique data that will help optimize strategic decisions and understand drug exposure at the site of action and the impact to the microenvironment. Adopting spatial bioanalysis and biology into your strategy early on, can help you save money and time in the long run.
Read More
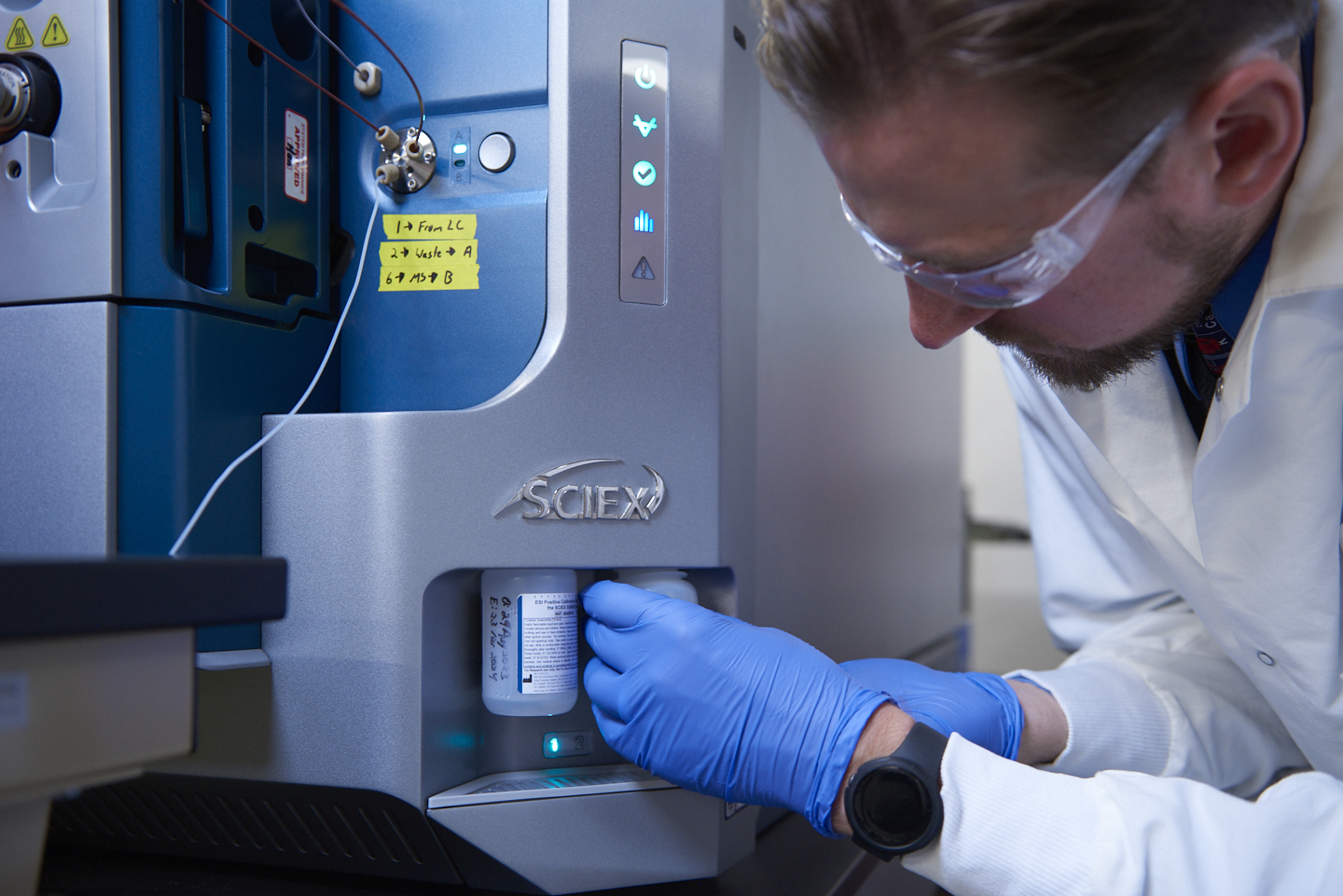

Aliri has the proven operational excellence across a range of traditional and cutting-edge technologies paired with the experience to support the unique needs of your development program.
Learn how you can utilize our team of dedicated scientists, program managers, quality specialists and industry experts.
Access quality data for filing your IND, NDA, or CTA with speed
Anticipate technical issues or potential roadblocks that may cause delays
Elucidate drug efficacy in the pertinent spatial context up to a cellular level, for more efficient and effective drug candidate evaluation and selection
Maximize the value of your drug discovery and development investments
131
Audits
16
FDA Audits
5